
DYOGEN group
web-code version: 2021-08-15
database version: 110.01
Tweets by GenomicusDB
 Contact us.
Contact us.

|

|
|

|

|

|


 Contact us.
Contact us.

|

|
|

|

|

|
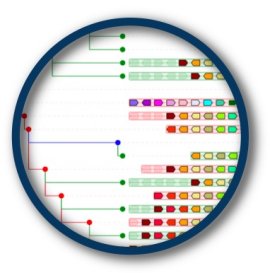
| Root species for S100Z (ENSPCOG00000005734) (Coquerels sifaka) | AlignView depth | PhyloView depth | |
|---|---|---|---|
| Fungi/Metazoa group |  ~1500 Mya ~1500 Mya |
399 species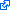  |
|
| Bilateria |  ~580 Mya ~580 Mya |
397 species  |
|
| Chordata |  ~550 Mya ~550 Mya |
393 species  |
|
| Vertebrata |  ~550 Mya ~550 Mya |
389 species  |
420 homologs (oldest homol.)  |
| Gnathostomata |  ~473 Mya ~473 Mya |
385 species  |
417 homologs  |
| Euteleostomi |  ~420 Mya ~420 Mya |
383 species  |
413 homologs  |
| Sarcopterygii |  ~400 Mya ~400 Mya |
253 species  |
222 homologs  |
| Tetrapoda |  ~359 Mya ~359 Mya |
251 species  |
218 homologs  |
| Amniota |  ~326 Mya ~326 Mya |
247 species  |
214 homologs  |
| Mammalia |  ~184 Mya ~184 Mya |
193 species  |
161 homologs  |
| Theria |  ~166 Mya ~166 Mya |
191 species  |
159 homologs  |
| Eutheria |  ~102 Mya ~102 Mya |
181 species  |
149 homologs  |
| Boreoeutheria |  ~95 Mya ~95 Mya |
171 species  |
139 homologs  |
| Euarchontoglires |  ~90 Mya ~90 Mya |
95 species  |
78 homologs  |
| Primates |  ~83 Mya ~83 Mya |
47 species  |
30 homologs  |
| Strepsirrhini |  ~69 Mya ~69 Mya |
7 species  |
7 homologs  |
| Lemuriformes |  ~37 Mya ~37 Mya |
5 species  |
5 homologs  |
| Lemuriformes_a |  ~37 Mya ~37 Mya |
3 species  |
3 homologs  |
| Coquerels sifaka |  ~0 Mya ~0 Mya |
1 species  |
1 homolog  |
-evalue <Real>
Expectation value (E) threshold for saving hits
Default = '10'
-word_size <Integer, >=2>
Word size for wordfinder algorithm
Default = '3'
-gapopen <Integer>
Cost to open a gap
Default = '11'
-gapextend <Integer>
Cost to extend a gap
Default = '1'
-matrix <String>
Scoring matrix name (normally BLOSUM62)
-threshold <Real, >=0>
Minimum word score such that the word is added to the BLAST lookup table
Default = '11'
-window_size <Integer, >=0>
Multiple hits window size, use 0 to specify 1-hit algorithm
Default = '40'